Crystal Structure ProGP
Allows user to search for known crystal structures (retrieved from PDB) of prokaryotic glycoproteins compiled under ProGP.
| ProGP ID | Organism Name | Protein Name | UniProtKB/SwissProt ID | PDB ID | Crystal Structure(PDB) |
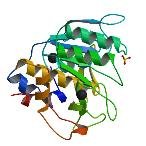
| ProGP197 (Subtilisin (SBL)-Cys mutant) | Bacillus lentus | Subtilisin (SBL)- Cys mutant | P29600 | 1JEA |
 |
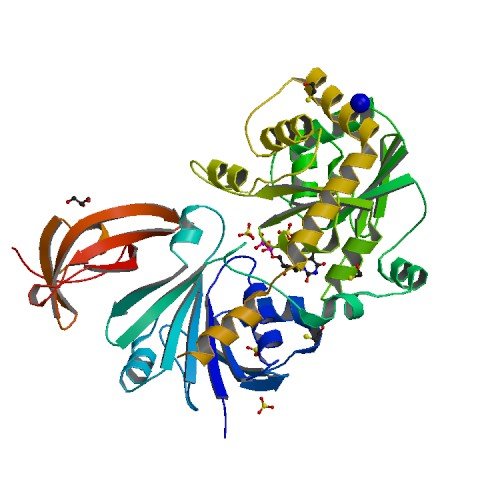
| ProGP542 (Translation elongation factor P (EF-P)) | Neisseria meningitidis HT1125 | Translation elongation factor P(EF-P) | E6MVW0 | 5WXK |
 |
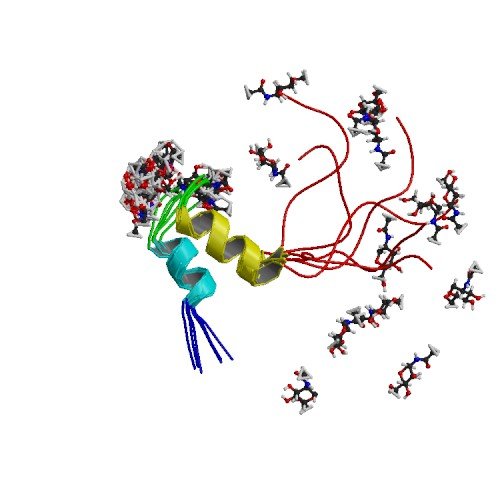
| ProGP402 (Glycocin F) | Lactobacillus plantarum KW30 | Glycocin F | E9K9Z1 | 2KUY |
 |
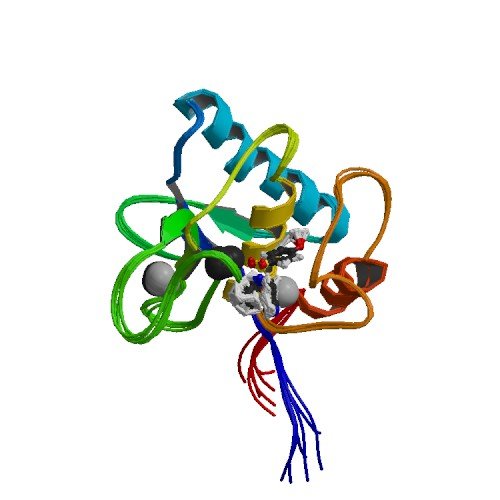
| ProGP211 (TMBP (Trehalose/maltose binding protein)) | Pyrococcus furiosus DSM 3638 | TMBP (Trehalose/maltose binding protein) | O51923 | 1EUB |
 |
| ProGP119 (Fimbrial protein (pilin)) | Neisseria gonorrhoeae N400 /MS11 | Fimbrial protein (pilin) | P02974 | 2HI2 2HIL 2PIL 1AY2 |
 .jpg) .jpg) .jpg) |
| ProGP4 (F-pilin) | Escherichia coli K12 | F-pilin | P04737 | 5LER 5LFB |
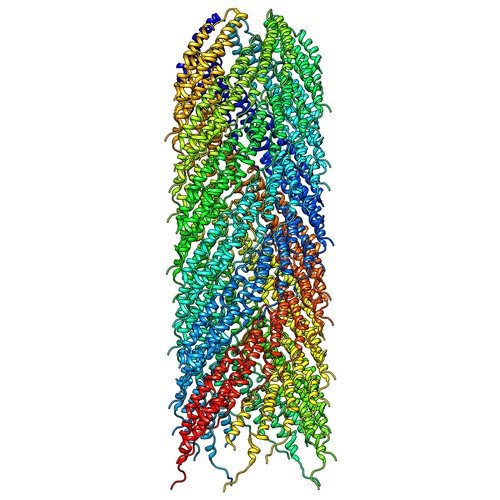 .jpg) |
| ProGP83 (Ice nucleation lipoglycoprotein) | Pseudomonas syringae strain S203 | Ice nucleation lipoglycoprotein | P06620 | 1INA 1INB |
 |
| ProGP282 (prG) | Mycobacterium tuberculosis | LprG | P0A5I8 | 3MH8 3MH9 3MHA |
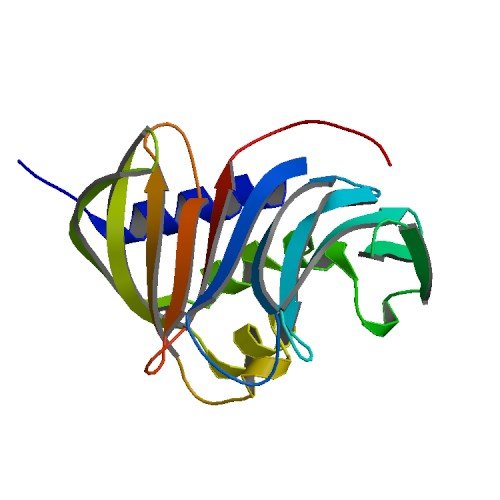 .jpg) .jpg) |
| ProGP347 (Spermidine/putrescine-binding protein (PotD)) | Escherichia coli K12/DH5alpha | Spermidine/putrescine-binding protein (PotD) | P0AFK9 | 1POT 1POY |
 .jpg) |
| ProGP101 (H(A16-M) (1,3-1,4)-beta-glucanase) | Paenibacillus (Bacillus) macerans and Bacillus amyloliquefaciens (velezensis) | H(A16-M) (1,3-1,4)-beta-glucanase | P23904 | 1CPM 1GLH 2AYH |
 .jpg) .jpg) |
| ProGP84 (Auracyanin-B1 (22 kDa)) | Chloroflexus aurantiacus | Auracyanin-B1 (22 kDa) | P27197 | 1OV8 1QHQ |
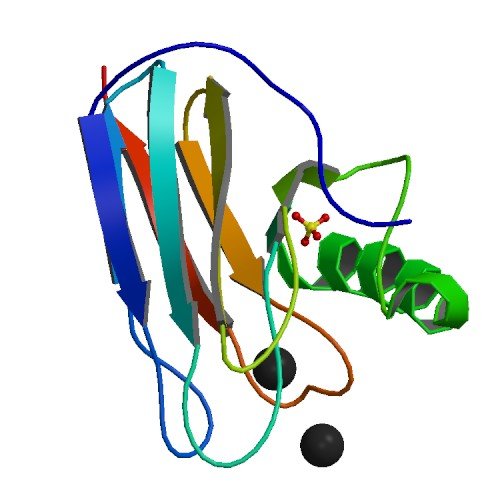 .jpg) |
| ProGP85 (Auracyanin-B2 (18 kDa)) | Chloroflexus aurantiacus | Auracyanin-B2 (18 kDa) | P27197 | 1OV8 1QHQ |
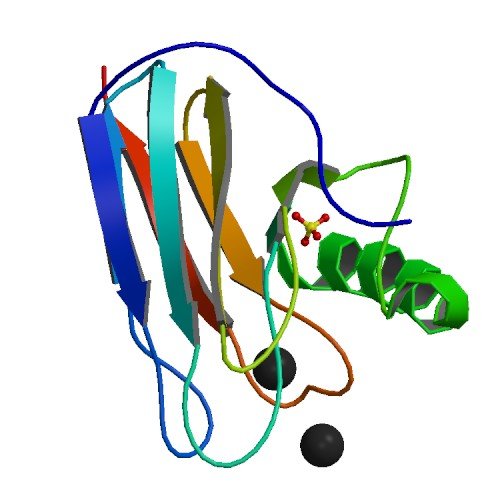 .jpg) |
| ProGP116 (Endo-β-N-acetylglucosaminidase F3) | Flavobacterium meningosepticum (Elizabethkingia meningoseptica) | Endo-β-N-acetylglucosaminidase F3 | P36913 | 1EOK 1EOM |
 .jpg) |
| ProGP212 (MDBP (Maltodextrin binding protein)) | Pyrococcus furiosus DSM 3638 | MDBP (Maltodextrin binding protein) | P58300 | 1ELJ |
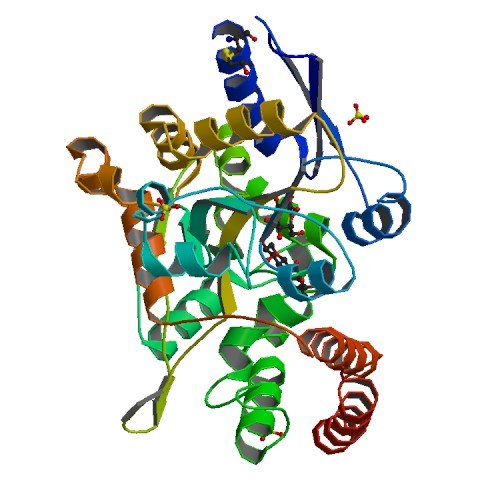 |
| ProGP346 (Chondroitinase ABC) | Proteus vulgaris | Chondroitinase ABC | P59807 | 1HN0 |
 |
| ProGP400 (Sublancin) | Bacillus subtilis 168 | Sublancin | P68577 | 2MIJ |
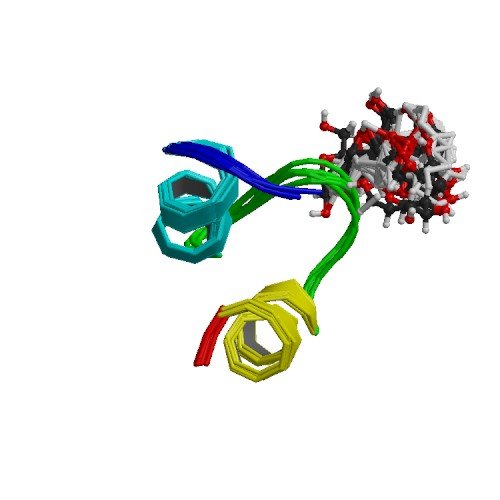 |
| ProGP189 (Superoxide dismutase [Cu-Zn]) | Mycobacterium tuberculosis H37Rv | Superoxide dismutase [Cu-Zn] | P9WGE9 | 1PZS |
 |
| ProGP273 (LppX (Putative lipoprotein)) | Mycobacterium tuberculosis | LppX (Putative lipoprotein) | P9WK65 | 2BYO |
 |
| ProGP270 (BlaC) | Mycobacterium tuberculosis | BlaC | P9WKD3 | 2GDN 3CG5 3IQA 3M6B 3M6H 3N8S 3NY4 |
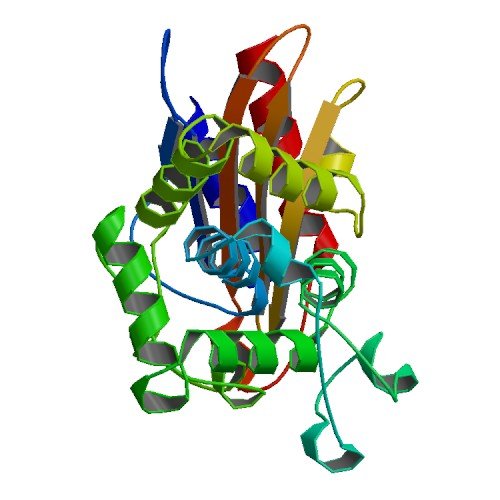 .jpg) .jpg) .jpg) .jpg) .jpg) .jpg) |
| ProGP384 (Cj0982c (Putative amino acid transporter periplasmic solute-binding protein CjaA)) | Campylobacter jejuni HB93-13 | Cj0982c (Putative amino acid transporter periplasmic solute-binding protein CjaA) | Q0P9S0 | 1XT8 |
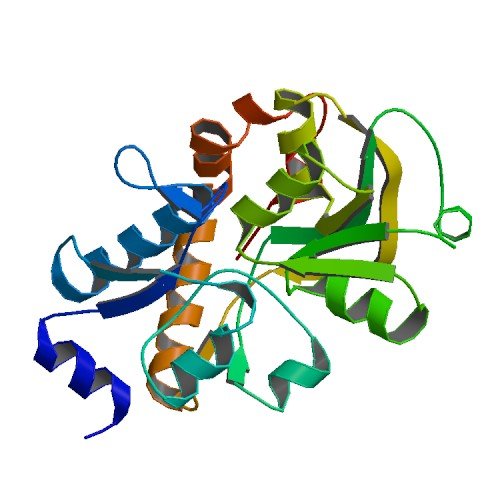 |
| ProGP222 (PEB3) | Campylobacter jejuni NCTC 11168 serotype O:2 | PEB3 | Q0PBL7 | 2HXW |
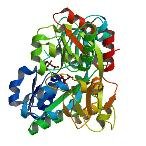 |
| ProGP166 (Chondroitinase-B) | Pedobacter heparinus (strain ATCC 13125 / DSM 2366 / NCIB 9290) | Chondroitinase-B | Q46079 | 1DBG 1DBO 1OFL 1OFM |
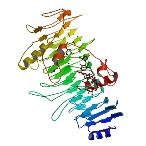 .jpg) .jpg) .jpg) |
| ProGP133 (Heparinase II) | Pedobacter heparinus (Flavobacterium heparinum) | Heparinase II | Q46080 | 2FUQ 2FUT |
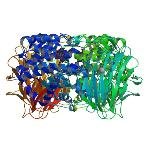 .jpg) |
| ProGP227 (HMW1 (Adhesin)) | Haemophilus influenzae 12 | HMW1 (Adhesin) | Q48031 | 2ODL |
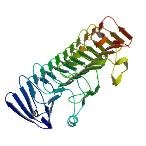 |
| ProGP108 (Tetrabrachion (S layer glycoprotein)) | Staphylothermus marinus F1 | Tetrabrachion (S-layer glycoprotein) | Q54436 | 1FE6 1YBK |
 .jpg) |
| ProGP132 (Chondroitinase-AC) | Pedobacter heparinus (strain ATCC 13125 / DSM 2366 / NCIB 9290) | Chondroitinase-AC | Q59288 | 1CB8 1HM2 1HM3 1HMU 1HMW |
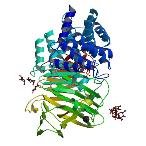 .jpg) .jpg) .jpg) .jpg) |
| ProGP216 (BclA (collagen-like protein)) | Bacillus anthracis str. Ames (Sterne 7702) | BclA (collagen-like protein) | Q81JD7 | 2R6Q |
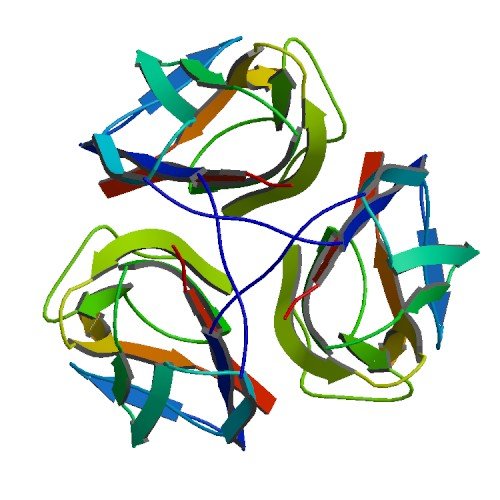 |
| ProGP510 (Translation elongation factor P (EF-P)) | Pseudomonas aeruginosa PAO1 | Translation elongation factor P(EF-P) | Q9HZZ2 | 3OYY |
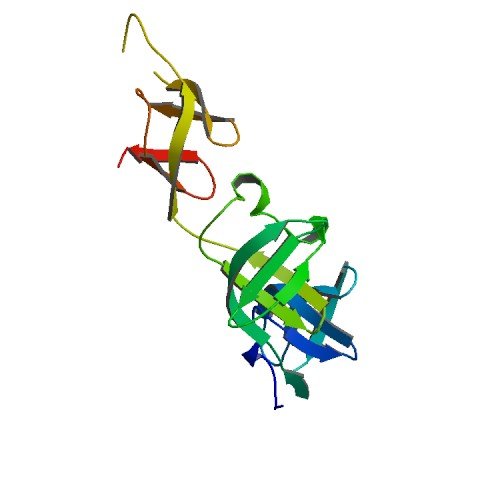 |
| ProGP171 (Man26A (b-mannosidase 26A)) | Cellulomonas fimi | Man26A (b-mannosidase 26A) | Q9XCV5 | 2BVT 2BVY 2X2Y |
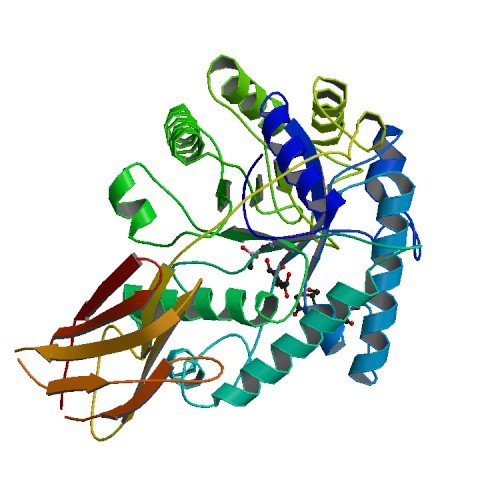 .jpg) .jpg) |
| ProGP209 (Fap1 (Fimbrial adhesin)) | Streptococcus parasanguis FW213 | Fap1 (Fimbrial adhesin) | Q9ZFF9 | 2KUB 2X12 |
 .jpg) |
| ProGP255 (HmcA) | Desulfovibrio gigas | HmcA | T2G9Q2 | 1Z1N |
 |
www.rcsb.org
H.M. Berman, J. Westbrook, Z. Feng, G. Gilliland, T.N. Bhat, H. Weissig, I.N. Shindyalov, P.E. Bourne. (2000) The Protein Data Bank Nucleic Acids Research, 28: 235-242.
H.M. Berman, J. Westbrook, Z. Feng, G. Gilliland, T.N. Bhat, H. Weissig, I.N. Shindyalov, P.E. Bourne. (2000) The Protein Data Bank Nucleic Acids Research, 28: 235-242.

